![PDF] ChIP and Re-ChIP Assays: Investigating Interactions Between Regulatory Proteins, Histone Modifications, and the DNA Sequences to Which They Bind PDF] ChIP and Re-ChIP Assays: Investigating Interactions Between Regulatory Proteins, Histone Modifications, and the DNA Sequences to Which They Bind](https://www.researchgate.net/profile/Susanna-Greer/publication/51825097/figure/fig2/AS:669356652494848@1536598470029/Re-ChIP-protocol-as-described-in-the-text-various-positive-and-negative-controls-for_Q320.jpg)
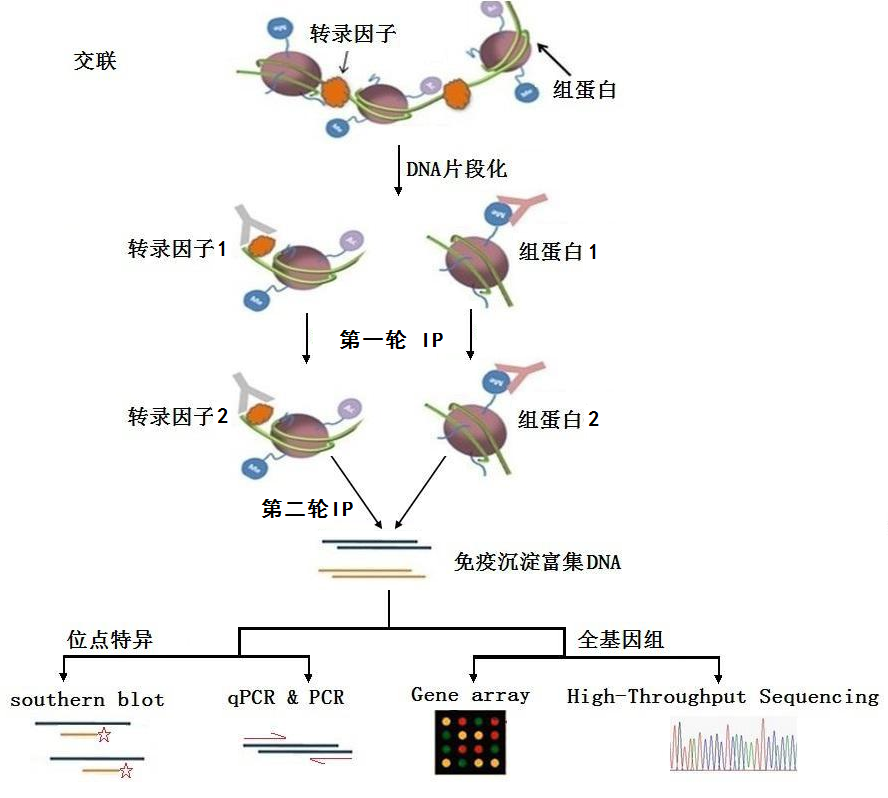
PDF] ChIP and Re-ChIP Assays: Investigating Interactions Between Regulatory Proteins, Histone Modifications, and the DNA Sequences to Which They Bind

reChIP-seq reveals widespread bivalency of H3K4me3 and H3K27me3 in CD4+ memory T cells | Nature Communications

ChIP and Re-ChIP Assays: Investigating Interactions Between Regulatory Proteins, Histone Modifications, and the DNA Sequences to Which They Bind | SpringerLink
The IKAROS Interaction with a Complex Including Chromatin Remodeling and Transcription Elongation Activities Is Required for Hematopoiesis | PLOS Genetics
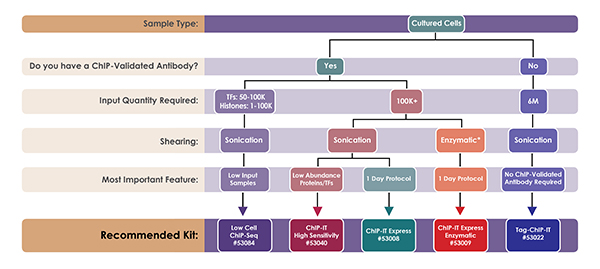
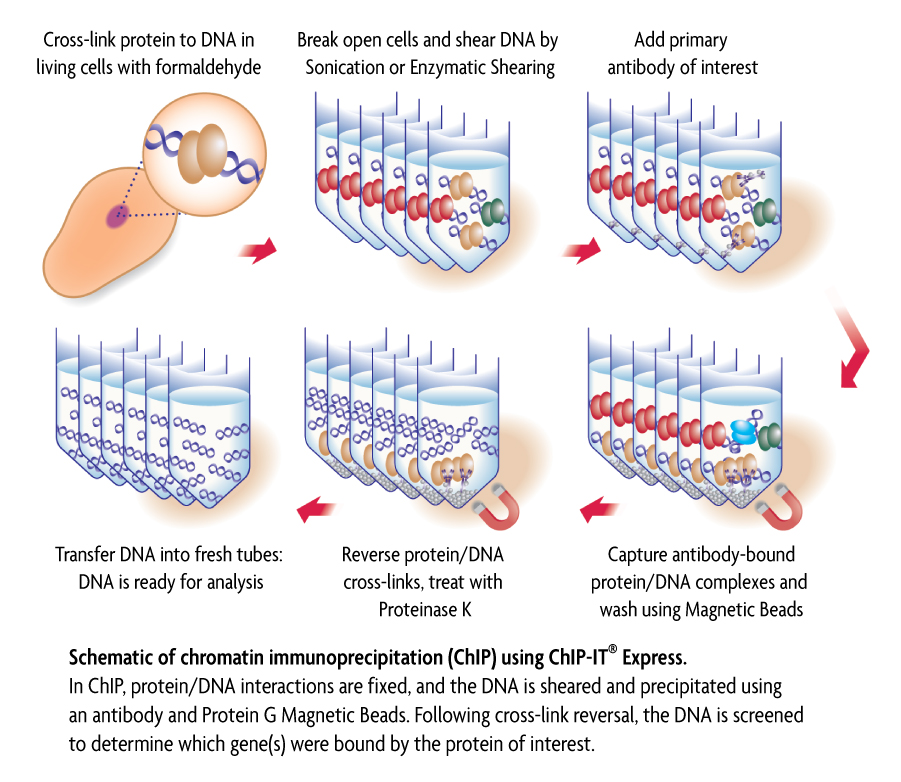
ChIP and Other Methods to Study DNA-Protein Interactions | Biocompare: The Buyer's Guide for Life Scientists
Enhancer-mediated enrichment of interacting JMJD3–DDX21 to ENPP2 locus prevents R-loop formation and promotes transcription

Bone-Specific Transcription Factor Runx2 Interacts with the 1α,25-Dihydroxyvitamin D3 Receptor To Up-Regulate Rat Osteocalcin Gene Expression in Osteoblastic Cells | Molecular and Cellular Biology

A parallelized, automated platform enabling individual or sequential ChIP of histone marks and transcription factors | PNAS
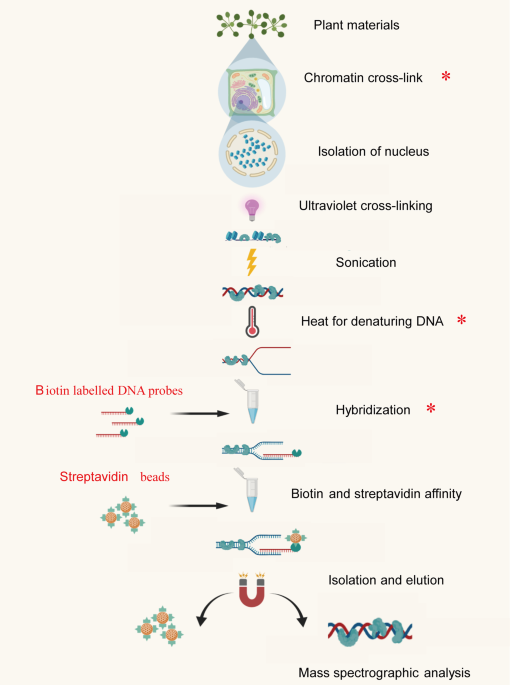
Reverse Chromatin Immunoprecipitation (R-ChIP) enables investigation of the upstream regulators of plant genes | Communications Biology






:max_bytes(150000):strip_icc()/10813-best-chocolate-chip-cookies-mfs-step-7-148-52cdaefcd6e04707863288ded8451075.jpg)